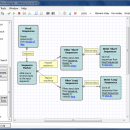
Unipro UGENE for Linux 50.0 freeware
Unipro UGENE for Linux is a free visual software solution for DNA and protein sequence analysis. UGENE provides customizable tools for visualization, analysis, annotation of genetic sequences. Key features are: Integrated MUSCLE alignment tool, Integrated HMMER2 package, OpenGL viewer for PDB macromolecular structures, and Workflow Designer to create and execute custom computational workflows. ...
| Author | Unipro |
| Released | 2024-04-14 |
| Filesize | 473.00 MB |
| Downloads | 2544 |
| OS | Linux |
| Installation | Instal And Uninstall |
| Keywords | DNA, education, visual software, analyze, genetic |
| Users' rating (59 rating) |
Unipro UGENE for Linux Free Download - we do not host any Unipro UGENE for Linux torrent files or links of Unipro UGENE for Linux on rapidshare.com, depositfiles.com, megaupload.com etc. All Unipro UGENE for Linux download links are direct Unipro UGENE for Linux download from publisher site or their selected mirrors.
| 50.0 | Apr 14, 2024 | New Release | Primer3 can be run without a target sequence. Primer-BLAST has been implemented in UGENE. The hairpin visualization tool mfold has been integrated into UGENE. The PlasMapper plasmids annotation database has been updated to the latest version. Python, included in the UGENE package, has been upgraded to version 3. The list of presets for Primer3 has been expanded. Improved compatibility with theme settings in macOS. Fixed issues related to the stability and usability of UGENE. |
| 49.1 | Nov 26, 2023 | New Release | Fix crash in genome aligner UI. |
| 49.0 | Nov 8, 2023 | New Release | The "Presets" feature has been developed for Primer3, that allows you to select a set of preset settings to run the tool. At the moment, two presets are available - default settings and Recombinase Polymerase Amplification. Primer3 has received the function of filtering the resulting primer pairs depending on their homo/heterocomplementarity. Annotation selection has been corrected in the Sequence Editor. Various minor interface fixes and enhancements. |